Electrostatics and ions in biology
 While it is well known that precise ionic conditions are required for stability and function of biomolecules, in particular RNA, quantifying their specific roles has been particularly challenging. One reason for this is that they have many modes by which they can impart stability. When they bind stably, they may be directly probed by structural biology methods (X-ray, cryo-EM, NMR). Even though these strongly-bound ions are very important for biological function, a larger number of ions associate more transiently. These transient ions can be associated with short-range (unidentate ‘chelated’), intermediate-range (‘diffuse’) and long-range (‘bulk’) interactions.
While it is well known that precise ionic conditions are required for stability and function of biomolecules, in particular RNA, quantifying their specific roles has been particularly challenging. One reason for this is that they have many modes by which they can impart stability. When they bind stably, they may be directly probed by structural biology methods (X-ray, cryo-EM, NMR). Even though these strongly-bound ions are very important for biological function, a larger number of ions associate more transiently. These transient ions can be associated with short-range (unidentate ‘chelated’), intermediate-range (‘diffuse’) and long-range (‘bulk’) interactions.
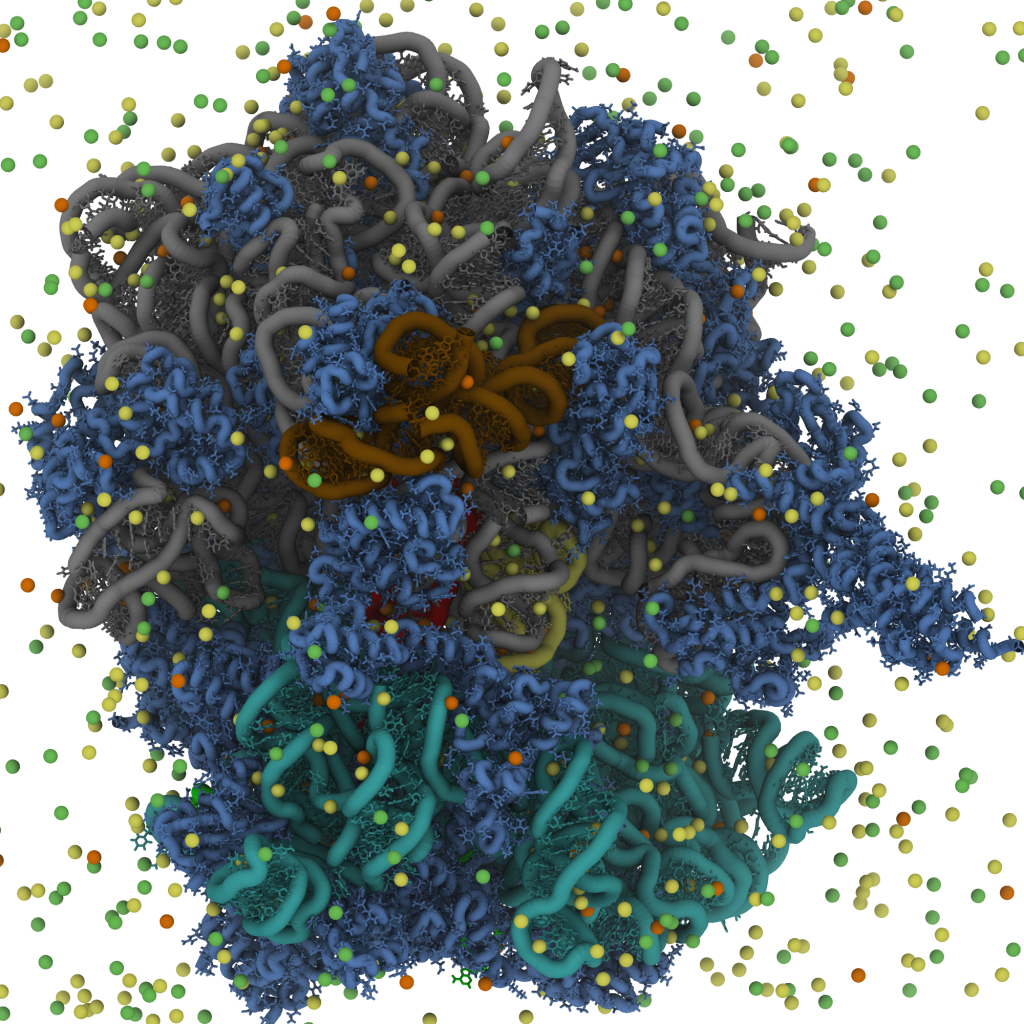
In the study of ions, we first developed an explicit-ion model that includes diffuse Mg2+, Cl- and K+ ions (Wang et al, JACS, 2022). Using this model, we were able to perform initial simulations of tRNA accommodation on the ribosome. This allowed us to quantify the concentration dependence of the free-energy landscape of tRNA on the ribosome, while also identifying specific ion-mediated interactions that contribute to the kinetics. To show the potential for these approaches, we also performed proof-of-principle simulations of the HIV-1 capsid with explicit ions (right), where we were able to describe the dynamics of ions around the capsid shell.
More recently, we extended our model, in order to account for short-range ‘inner-shell’ interactions (Wanes et al, PNAS, 2026). With this new model, we tackled a larger-scale motion: subunit rotation. In the eukaryotic ribosome, subunit rotation involves the collective displacement of approximately 80k atoms, relative to roughly 120k atoms. By using our SMOG-based strategies for simulating conformational motions in biomolecules, along with this explicit-ion model, we were able to show how the free-energy landscape of rotation responds to mM changes in ionic concentrations. This showed how a combination of inner-shell and outer-shell interactions bridge the subunits, in a concentration-dependent manner. We anticipate that this model will continue to be an effective tool for isolating the precise contributions of monovalent and divalent ions to large-scale biomolecular dynamics.
